INTRODUCTION
Histone deacetylase 10 (HDAC10) has been implicated in several pathological processes, including cancer progression and inflammatory disease [1–4]. In cervical cancer, HDAC10 suppresses cell invasion and metastasis by binding to matrix metalloproteinase 2 (MMP2) and matrix metalloproteinase 9 (MMP9) promoters and deacetylating histones, thereby reducing the expression of these MMPs [1]. In allergic airway inflammation, HDAC10 promotes macrophage M2 polarization by deacetylating signal transducer and activator of transcription 3 (STAT3), indicating that it can directly act on transcription factors and chromatin [3]. These findings suggest that HDAC10 participates in disease-relevant signaling pathways through the deacetylation of specific nuclear substrates and raise the question of which additional nuclear proteins are targeted by HDAC10 in such contexts.
The activator protein-1 (AP-1) component c-Jun is a key regulator of genes involved in proliferation, stress responses, inflammation, and tumorigenesis. It controls the expression of MMPs and inflammatory mediators that are also influenced by HDAC10 [5–8]. The activity of c-Jun is regulated not only by c-Jun N-terminal kinase (JNK)-mediated phosphorylation but also by reversible lysine acetylation, which modulates its DNA-binding properties and transcriptional output at AP-1-responsive promoters [6, 7, 9]. Coactivators like p300/CBP can acetylate lysine residues in the basic region of c-Jun. Moreover, changes in acetylation at these sites alter promoter selectivity and the balance between activation and repression, indicating that acetylation fine-tunes AP-1 signaling instead of acting as a simple on/off switch [9]. Thus, deacetylases that directly remove these modifications are expected to have a crucial impact on c-Jun-dependent gene regulation [10].
Several deacetylases have been reported to influence the AP-1/c-Jun axis [11, 12]. HDAC3 binds to the N-terminus of c-Jun and represses AP-1-dependent transactivation. It can also deacetylate histones at the c-Jun promoter to reduce c-Jun expression [13]. SIRT1 interacts with c-Jun, inhibits AP-1/MMP9 transcriptional activity, and modulates inflammatory gene expression by deacetylating c-Jun or nearby histones as a nonclassical, NAD+-dependent deacetylase [14, 15]. Other HDACs, such as HDAC1, HDAC2, HDAC4, and HDAC6, have mainly been linked to chromatin-level regulation of c-Jun transcription or to upstream signaling pathways that converge on c-Jun [16, 17]. However, despite these observations, the deacetylases that directly recognize c-Jun as a nuclear substrate and the functional impacts of this interaction on AP-1-dependent transcription remain insufficiently understood [11, 18].
Considering the emerging role of HDAC10 as a nuclear deacetylase for STAT3 and its involvement in pathways that overlap with c-Jun-regulated gene expression, we hypothesized that c-Jun represents a previously unrecognized nuclear substrate of HDAC10 [3, 9]. In this study, we examined whether HDAC10 associates with c-Jun, whether it deacetylates c-Jun and a δ-domain deletion mutant (Δδ c-Jun), and how it affects AP-1-dependent transcription and c-Jun DNA-binding activity in human embryonic kidney 293 (HEK293) cells [9].
MATERIALS AND METHODS
HEK293 cells were maintained in Dulbecco’s modified Eagle’s medium (high glucose) supplemented with 10% heat-inactivated fetal bovine serum and 100 U/mL penicillin–streptomycin (all from HyClone, Logan, UT, USA) at 37℃ in a humidified incubator with 5% CO₂. Cells were transiently transfected with the indicated expression plasmids using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol, and analyzed 24–36 hr post-transfection.
Human c-Jun was expressed as an xpress-tagged construct in a mammalian expression vector. Δδ c-Jun, in which the N-terminal δ-domain is deleted, and the Ser63/Ser73 phosphorylation-defective mutant were generated by PCR-based mutagenesis and verified by DNA sequencing, as described previously for Jun mutants. HDAC10 was expressed as an N-terminal FLAG-tagged protein. Catalytic-domain mutants or truncations of HDAC10 were generated by standard PCR cloning and sequencing. AP-1-luciferase and Renilla luciferase (internal control) reporter plasmids were obtained from Promega (Madison, WI, USA).
For co-immunoprecipitation, transfected HEK293 cells were lysed in NP-40 lysis buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% NP-40, 1 mM EDTA) containing Protease and Phosphatase Inhibitor Cocktail (Sigma-Aldrich, St. Louis, MO, USA). Lysates were cleared by centrifugation and incubated with anti-FLAG or anti-xpress antibody for 2 hr at 4℃, followed by incubation with protein A/G agarose beads for an additional 1–2 hr. Beads were washed three to four times with lysis buffer and bound proteins were eluted by boiling in SDS sample buffer. Samples were resolved by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and analyzed by immunoblotting with antibodies against c-Jun, FLAG, xpress, acetyl-lysine, HDAC10, and β-actin (all antibodies were purchased from Santa Cruz Biotechnology, Dallas, TX, USA). Immunoreactive bands were visualized by enhanced chemiluminescence.
To assess c-Jun acetylation, xpress-tagged c-Jun or Δδ c-Jun was immunoprecipitated from transfected HEK293 cell lysates using anti-xpress antibody as described above. Immunoprecipitates were subjected to SDS–PAGE and blotted with anti–acetyl-lysine antibody to detect acetylated c-Jun species, followed by reprobing with anti-c-Jun antibody to monitor total precipitated c-Jun.
AP-1 transcriptional activity was measured using a firefly luciferase reporter under the control of multimerized AP-1 binding sites. HEK293 cells in 24-well plates were co-transfected with AP-1-luciferase reporter, Renilla luciferase control plasmid, and the indicated combinations of c-Jun (or Δδ c-Jun) and HDAC10 expression vectors. At 24 hr after transfection, cells were lysed in passive lysis buffer and firefly and Renilla luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega) with a luminometer according to the manufacturer’s protocol. Firefly luciferase values were normalized to Renilla luciferase to control for transfection efficiency and are presented as relative fold induction over vector control.
c-Jun DNA-binding activity was quantified using the TFact™ c-Jun DNA-Binding enzyme-linked immunosorbent assay (ELISA) kit (Assay Biotechnology Company, Sunnyvale, CA, USA) following the manufacturer’s instructions. Briefly, nuclear extracts were prepared from HEK293 cells transfected with c-Jun in the presence or absence of HDAC10, using the extraction buffers supplied with the kit supplemented with PMSF and protease inhibitors. Equal amounts of nuclear protein were incubated in 96-well plates pre-coated with a double-stranded oligonucleotide containing the AP-1 (TRE) consensus sequence. After washing, bound c-Jun was detected with a specific primary antibody and HRP-conjugated secondary antibody provided in the kit, and color was developed with TMB substrate. Absorbance was read at 450 nm using a microplate reader, and values were expressed as relative DNA-binding activity normalized to the control condition.
All experiments were performed at least three times with independent cell preparations. Data are expressed as mean ± S.D. Statistical significance was evaluated using Student’s t-test or one-way ANOVA with appropriate post-hoc tests, and p values<0.05 were considered significant.
RESULTS
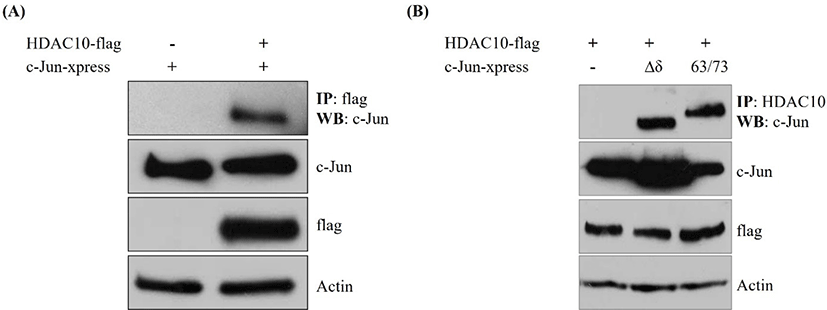
To determine whether HDAC10 associates with c-Jun, we performed co-immunoprecipitation in cells co-transfected with HDAC10-FLAG and xpress-tagged c-Jun. Immunoprecipitation of HDAC10 with an anti-FLAG antibody brought down c-Jun, whereas no c-Jun signal was detected in lysates lacking HDAC10 (Fig. 1A). Input blots revealed comparable levels of c-Jun, HDAC10, and actin between samples. We next assessed whether this interaction depends on the N-terminal δ-domain or phosphorylation at Ser63/Ser73 of c-Jun. Co-immunoprecipitation using Δδ c-Jun or a Ser63/Ser73 phosphorylation mutant revealed that both mutants were readily detected in HDAC10 immunoprecipitates (Fig. 1B). These findings indicate that HDAC10 binds to c-Jun independently of the δ-domain and the phosphorylation status of Ser63/Ser73.
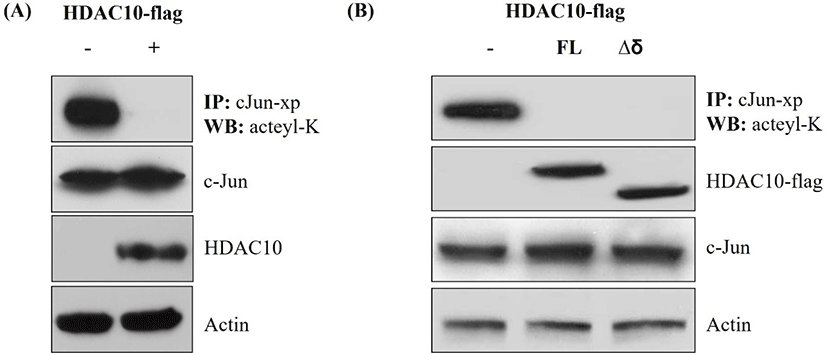
To determine whether HDAC10 regulates the acetylation status of c-Jun, we immunoprecipitated Xpress-tagged c-Jun and analyzed it by immunoblotting with an anti-acetyl-lysine antibody. Co-expression of HDAC10 markedly reduced acetyl-lysine signals on c-Jun, while total c-Jun levels in the input were similar between conditions (Fig. 2A). Moreover, HDAC10 and actin levels were comparable across samples. The same experiment was conducted using Δδ c-Jun. In cells expressing HDAC10, acetyl-lysine signals on Δδ c-Jun were decreased; however, input blots showed similar expression levels of Δδ c-Jun and HDAC10 relative to controls (Fig. 2B). Thus, HDAC10 deacetylates both full-length c-Jun and Δδ c-Jun, indicating that its deacetylase activity does not require the δ-domain.
We assessed the functional impact of HDAC10 on c-Jun-dependent transcription using an AP-1 luciferase reporter assay. The expression of c-Jun strongly increased AP-1-driven luciferase activity relative to the vector control (Fig. 3A). Co-expression of HDAC10 at 0.1, 0.5, and 1 µg reduced reporter activity in a dose-dependent manner. We then performed the same assay using Δδ c-Jun and found that Δδ c-Jun significantly enhanced AP-1 reporter activity. This increase was progressively attenuated by the co-expression of HDAC10 (Fig. 3B). These results indicate that HDAC10 represses AP-1-dependent transcription driven by both full-length c-Jun and Δδ c-Jun.
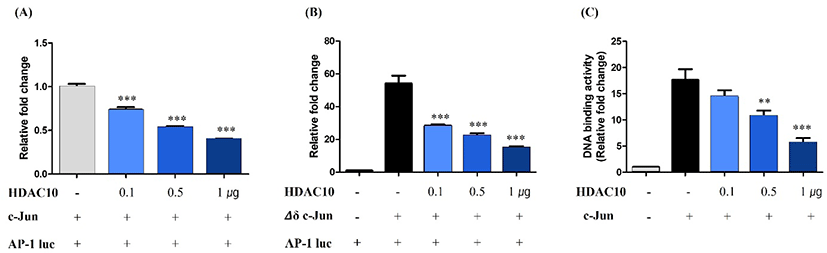
To further determine how HDAC10 inhibits c-Jun activity, we measured c-Jun DNA-binding activity using an ELISA-based AP-1 DNA-binding assay. Compared with the control cells, the expression of c-Jun increased binding to an AP-1 consensus oligonucleotide (Fig. 3C). However, co-expression of HDAC10 reduced c-Jun DNA-binding activity in a dose-dependent manner. Together, these findings indicate that HDAC10 negatively regulates c-Jun by deacetylating c-Jun and diminishing its DNA-binding capacity, thereby suppressing AP-1-dependent transcription.
DISCUSSION
HDAC10 is a class IIb deacetylase implicated in cancer progression and inflammatory disease, but the nuclear substrates through which it exerts these effects have not been fully identified [1, 2, 19]. In this study, we found that c-Jun is a nuclear substrate of HDAC10 and that HDAC10 negatively regulates AP-1-dependent transcription. HDAC10 formed a complex with c-Jun in HEK293 cells, reduced p300-induced lysine acetylation of wild-type c-Jun and Δδ c-Jun, suppressed c-Jun-driven AP-1 luciferase reporter activity, and reduced c-Jun DNA-binding activity. These effects were dependent on the catalytic activity of HDAC10 and occurred independently of the c-Jun δ-domain, suggesting that HDAC10 constrains c-Jun activity through the deacetylation of lysine residues in the DNA-binding or transactivation domains. These findings identify a mechanism by which HDAC10 modulates AP-1 signaling through regulation of c-Jun acetylation.
Previous studies have linked HDAC10 to the regulation of cancer cell invasion and inflammatory responses [1, 3]. In cervical cancer cells, HDAC10 binds to MMP2 and MMP9 promoters and deacetylates histones, thereby suppressing the transcription of these MMPs and inhibiting metastasis [1]. In macrophages, HDAC10 deacetylates STAT3 to promote M2 polarization and exacerbate allergic airway inflammation [3]. The findings of our study extend these observations by demonstrating that HDAC10 acts on the transcription factor c-Jun. As c-Jun regulates the expression of MMPs and inflammatory mediators, such as cytokines and chemokines [20–22], HDAC10-mediated deacetylation of c-Jun provides a plausible mechanism by which HDAC10 controls these gene networks in pathological contexts [9].
For example, in tumor cells, where HDAC10 expression is maintained or upregulated, the deacetylation of c-Jun may limit AP-1 activity and thereby restrain invasion. On the other hand, reduced HDAC10 activity could facilitate the hyperacetylation and activation of c-Jun, promoting metastatic progression [1]. Similarly, in inflammatory settings, HDAC10 may fine-tune the balance between pro- and anti-inflammatory gene expression through both STAT3 and c-Jun [3].
Our results place HDAC10 alongside other deacetylases known to influence the AP-1/c-Jun axis. HDAC3 binds to the N-terminus of c-Jun and represses AP-1 transactivation. It can also deacetylate histones at the c-Jun promoter to downregulate c-Jun expression; however, direct deacetylation of c-Jun by HDAC3 has not been mapped [13]. SIRT1 interacts with c-Jun, inhibits AP-1/MMP9 transcriptional activity, and modulates inflammatory gene expression in T cells by deacetylating c-Jun or adjacent histones [14]. Additional class I and class II HDACs, including HDAC1, HDAC2, HDAC4, and HDAC6, have been associated with chromatin modifications at AP-1 target loci or with upstream regulation of JNK–c-Jun signaling [16, 17, 23]. In comparison, HDAC10 is a distinct class IIb deacetylase that deacetylates c-Jun and suppresses its DNA-binding capacity and transcriptional output, independently of the δ-domain. Whether HDAC10, HDAC3, and SIRT1 target overlapping or distinct lysine residues on c-Jun, and how these modifications are coordinated with JNK phosphorylation or ubiquitination, will be important questions for future investigation [24, 25].
The identification of c-Jun as an HDAC10 substrate has potential implications for understanding the role of HDAC10 in cancer and inflammation. In tumors driven by hyperactive AP-1, such as those with upregulated c-Jun or JNK signaling, HDAC10 may act as a negative feedback regulator by limiting c-Jun acetylation and activity [26, 27]. Consistent with this finding, HDAC10 expression correlates with reduced metastatic potential in some cancers [1]. In contrast, loss of HDAC10 could unleash c-Jun activity and contribute to tumor progression, tumor invasion, or resistance to therapy. In inflammatory diseases, HDAC10-mediated deacetylation of c-Jun may modulate the expression of AP-1 target genes, such as those encoding cyclooxygenase-2 (COX-2) or MMPs, thereby influencing immune cell function or tissue remodeling [20, 28]. Pharmacological or genetic inhibition of HDAC10, which is already under consideration for cancer therapy, may therefore exhibit pleiotropic effects through both STAT3 and c-Jun pathways [2, 4].
In summary, our study identifies c-Jun as a non-histone substrate of HDAC10 and demonstrates that HDAC10 regulates c-Jun acetylation, thereby negatively modulating AP-1 signaling. These findings expand the known repertoire of non-histone substrates for HDAC10 and provide mechanistic insight into how HDAC10 influences AP-1 dependent gene expression. Although the present study primarily employed overexpression systems to examine the regulatory interaction between HDAC10 and c-Jun, complementary loss-of-function approaches will be valuable for further defining the endogenous role of HDAC10 in controlling c-Jun acetylation and AP-1 signaling. In addition, validation in disease-relevant cellular or physiological models will help clarify the biological significance of this regulatory axis. Together, these findings provide a foundation for future studies aimed at defining the in vivo roles of the HDAC10–c-Jun pathway and evaluating its potential therapeutic relevance.







